 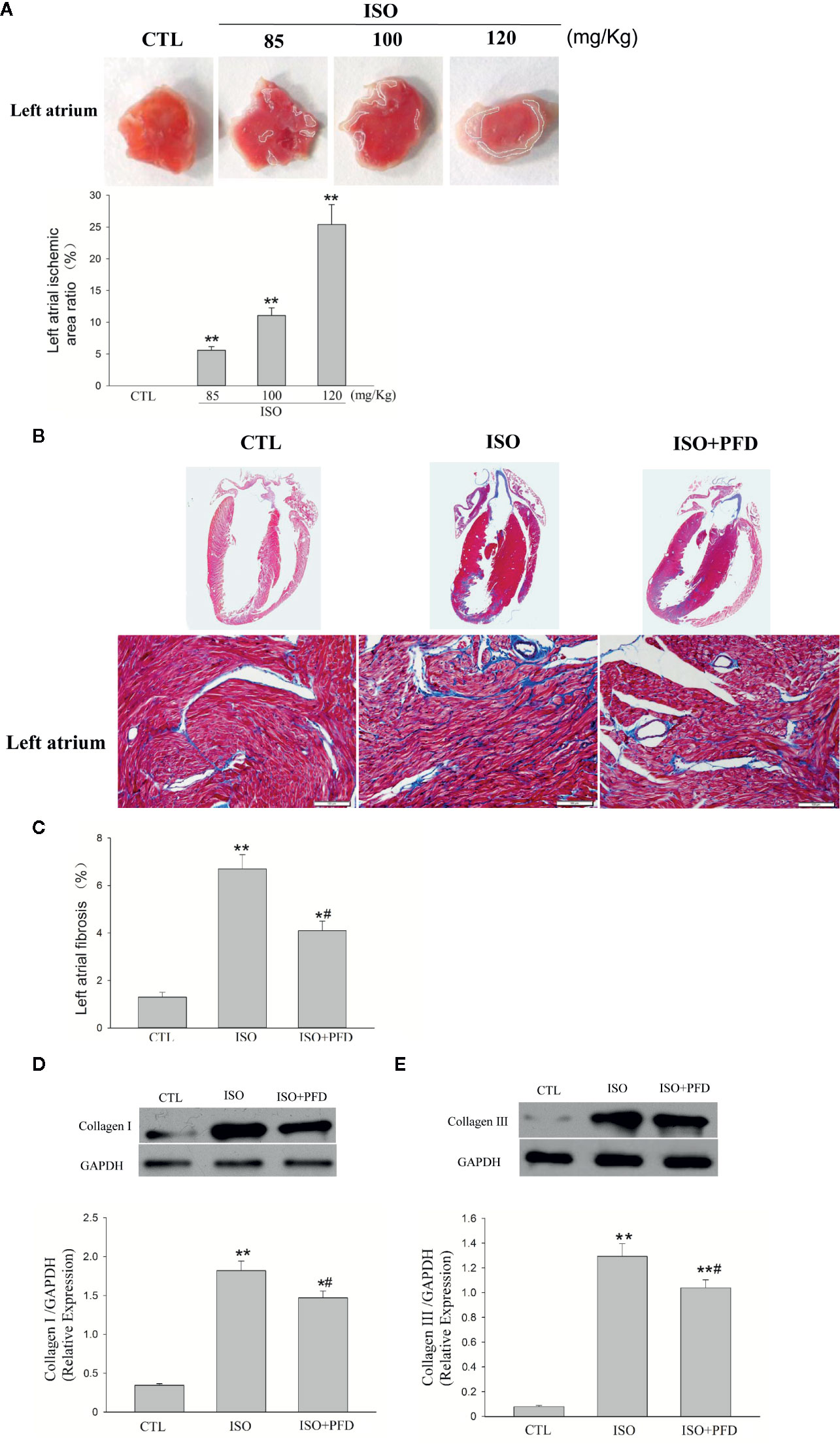
We recently had to perform a couple (always trying to please grant reviewers) and while I was analyzing the data using ImajeJ I started to wonder the exact same thing. I find this post extremely interesting, my lab is a physiology lab and we try not to do western blots unless we absolutely have to. Since your analysis says there is, it is important to know if this difference is biologically relevant. Again, this is all just me observing this band without knowing the scientific relevance (or really the experiment at all), but when I look at those bands I do not see a difference in intensities. This also brings to light a point that sometimes we rely to heavily on the "statistical significance" of these band intensities. Or if it would just be another layer of area selection bias. Me wonder if the loading control would help. Regardless your analysis could shows the consequences of these types of analyses, and it makes I realize this was just a quick experiment of intrigue, and it looks like maybe that top band is the control, or a spurious band. I am curious if you did the same analysis with a loading control, and then normalized the signal. Something like a technical replicate of the area selection? As you mentioned, it is important to correct for noise and that is another added step of variance and potential bias. It makes sense that there is variation between your attempts because of the area around the band, and I wonder if it would be possible to take that into consideration when you are doing western blot analysis. Coomassie staining as loading control in Western blot analysis.This is a really interesting analysis, that I have always wondered about. Proceedings of the National Academy of Sciences, 76(9), 4350–4354. Electrophoretic transfer of proteins from polyacrylamide gels to nitrocellulose sheets: procedure and some applications. A defined methodology for reliable quantification of Western blot data. Size matters: how sample size affects the reproducibility and specificity of gene set analysis. An error model for protein quantification. Quantifying western blots: none more black. Expert Review of Proteomics, 11, 549–560. The necessity of and strategies for improving confidence in the accuracy of western blots. Total Protein Analysis as a Reliable Loading Control for Quantitative Fluorescent Western Blotting. Misleading westerns: common quantification mistakes in western blot densitometry and proposed corrective measures. The use of total protein stains as loading controls: An alternative to high-abundance singleprotein controls in semiquantitative immunoblotting. Keywords: blotting, statistical analysis, non-parametric, densitometryĪldridge, G.M. Furthermore, repeating western blotting on a set of particular proteins may improve the analysis part as well. This procedure of quantifying western blotting is comprehensive, simple and can be applied to collect data in compliance with statistical norms. The differences among various blots that correspond to the expressed target proteins are tested viably using appropriate statistical tools. In present study, we attempt a noble approach while analyzing an image from western blotting using Gaussian blur as filter and thereby generating data in order to perform meaningful statistical analysis. 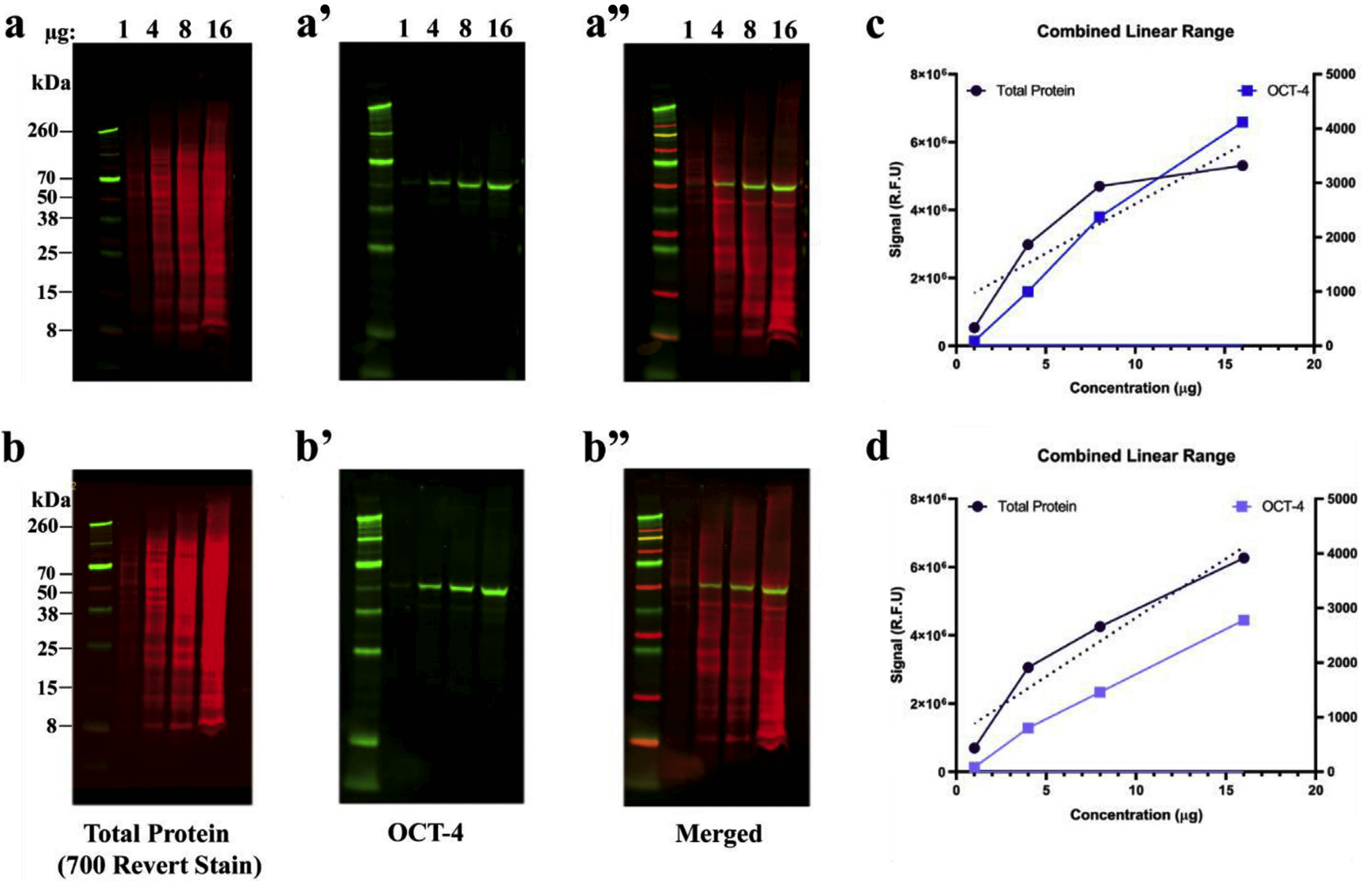
At the same time, extraction of the poor sample size (n=3/5/7) turns such analysis non-Gaussian and less robust to statistical errors. As a consequence, quantitative analysis of expression of protein through western blotting often suffers from data inconsistency. However, there is no unified method for quantifying the expression of proteins. In biological sciences, western blotting technique is widely used to quantify the expression of proteins in a given sample. Article Details: Received: | Accepted: | Available online: 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
